An Integrated Discovery Pipeline
The project follows a multidisciplinary and iterative workflow in which computational modeling, chemical synthesis, and biological evaluation are tightly interconnected.
Computational predictions guide compound design, experimental data validate and refine molecular hypotheses, and biological results inform further optimization. This continuous feedback loop enables the progressive identification of small molecules capable of modulating bacterial communication pathways.
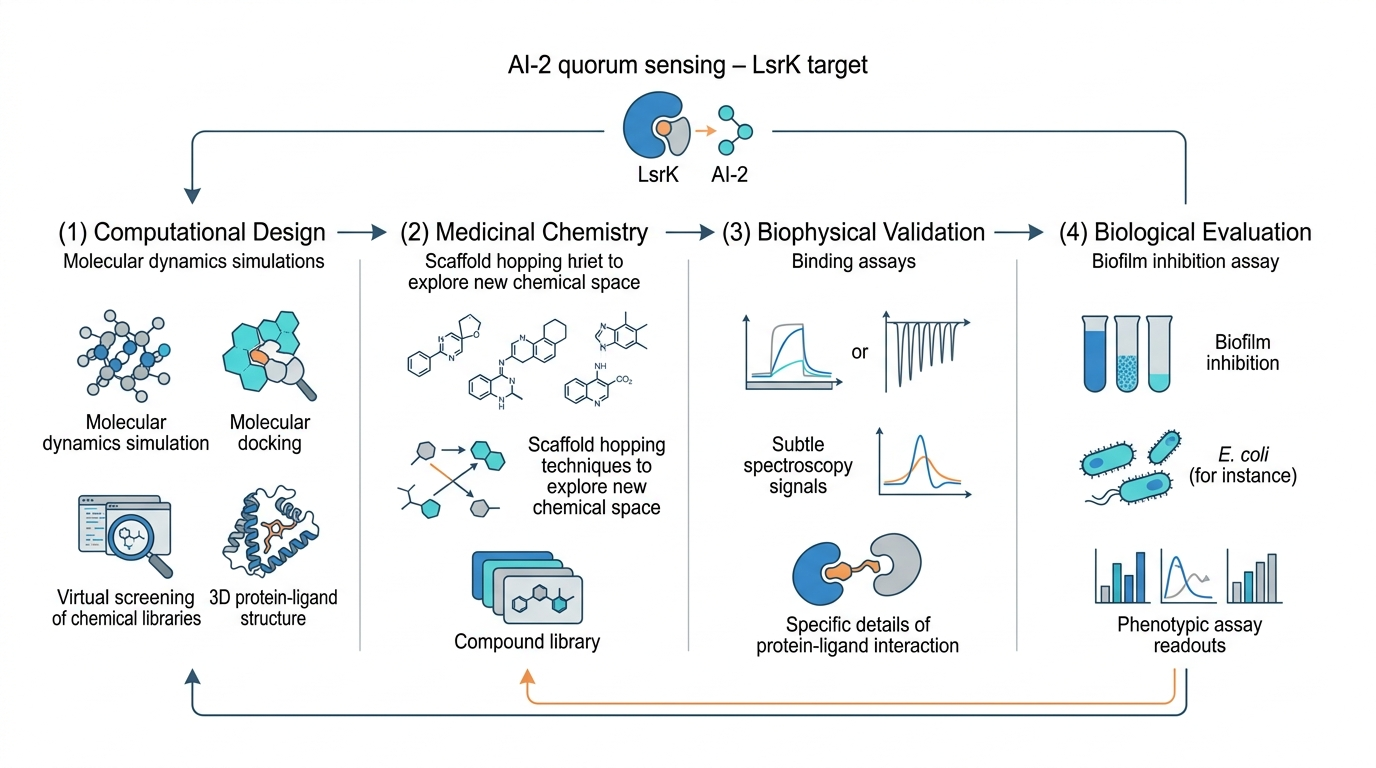 Multidisciplinary workflow for the discovery of LsrK-targeting antivirulence agents
Multidisciplinary workflow for the discovery of LsrK-targeting antivirulence agents
Iterative integration of computational modeling, medicinal chemistry, and biological validation
ACTIVITY 1 – Computational Design and Ligand Identification
The project integrates advanced computational approaches to support ligand discovery and rational design. Molecular dynamics simulations and structure-based virtual screening are employed to explore the conformational landscape of the target protein and to identify potential binding sites.
These strategies enable the selection of candidate ligands from large chemical libraries, including both novel chemotypes and approved drugs explored through repurposing approaches. Computational analyses also guide the prioritization of compounds for synthesis and provide structural hypotheses that drive the iterative optimization of ligand–target interactions .
ACTIVITY 2 – Medicinal Chemistry and Chemical Space Exploration
Based on computational insights, the project develops multiple libraries of small molecules targeting LsrK through an integrated medicinal chemistry strategy.
Different design approaches are explored in parallel, including hit optimization, scaffold diversification, and substrate-inspired design aimed at mimicking structural features of the natural AI-2 precursor. This strategy enables a broad exploration of the chemical space associated with LsrK recognition, leading to the generation of structurally diverse compound libraries suitable for biological evaluation.
Particular attention is devoted to the development of efficient and sustainable synthetic methodologies, allowing the rapid preparation and purification of compounds within an iterative design–synthesis–test cycle .
ACTIVITY 3 – Biophysical and Biochemical Characterization
The interaction between candidate molecules and the LsrK target is investigated through complementary biophysical techniques, including spectroscopic and NMR-based approaches.
These methods provide direct evidence of ligand binding and enable the characterization of the molecular determinants governing protein–ligand interactions. The integration of orthogonal experimental techniques allows the validation of computational predictions and supports the development of structure–activity relationships guiding further optimization .
ACTIVITY 4 – Evaluation of Antibiofilm and Antivirulence Activity
The biological activity of the synthesized compounds is assessed through microbiological assays designed to evaluate their impact on quorum sensing–regulated phenotypes.
In particular, compounds are tested for their ability to inhibit biofilm formation and to modulate virulence-related processes in representative Gram-positive and Gram-negative pathogens. The experimental workflow is specifically designed to distinguish antivirulence effects from antibacterial activity, ensuring that the identified molecules do not significantly affect bacterial growth.
This approach allows the identification of compounds capable of attenuating bacterial pathogenicity while preserving cell viability, in line with the antivirulence strategy of the project .
